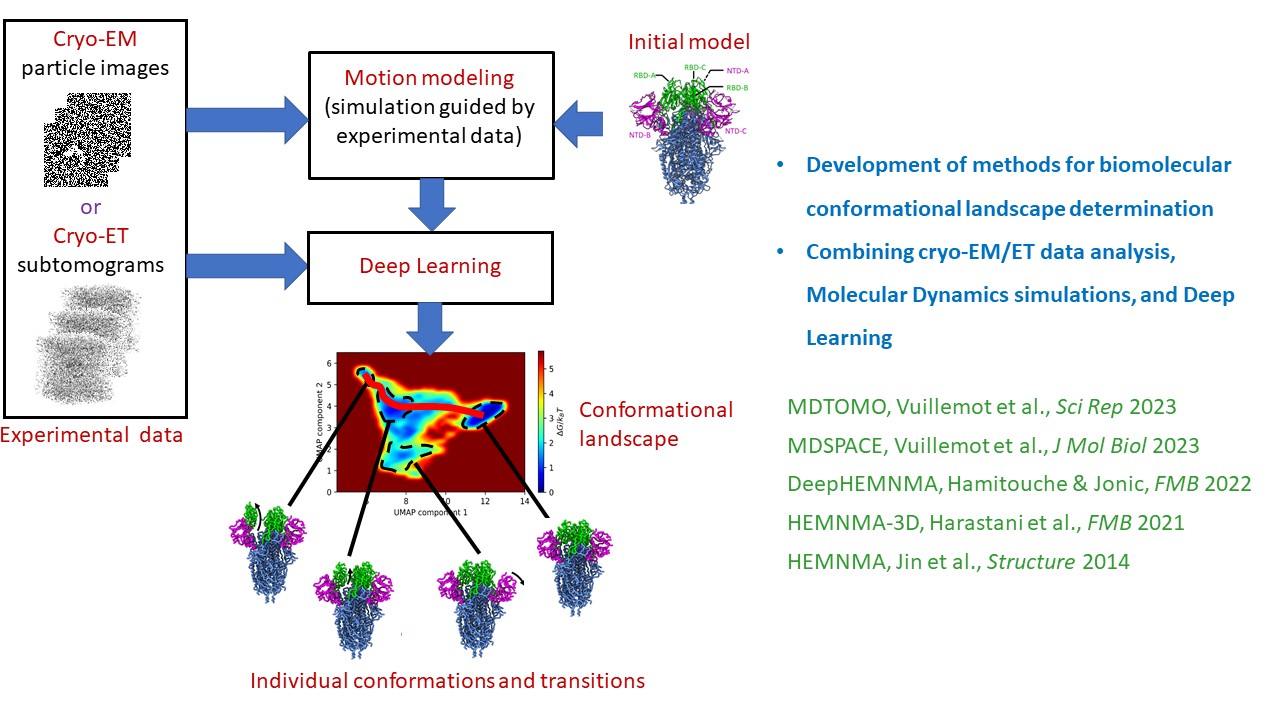
We develop algorithms and software that combine image analysis, molecular dynamics simulations, and artificial intelligence to enable full exploration of biomolecular conformational dynamics by in vitro and in situ cryo electron microscopy and cryo electron tomography. These methods are used to study biomedically important molecular complexes in collaboration with experimentalists. For highlights on this research, see below.
Alex Mirzaei, Postdoc (06/202-11/2021)
Jing Wang, Master intern (03/2024-08/2024), Sorbonne University, Paris (currently PhD student at Grenoble Alpes University)
Shun Robert, Master intern (03/2024-08/2024), Paris-Saclay University, Orsay
Guchao Zeng, Master intern (05/2023-10/2023), CentraleSupélec, Gif-sur-Yvette
Domique Rado Rakoto, Master intern (05/2023-10/2023), CY Tech, Cergy Pontoise
Nima Barati Moghadam, Master intern (07/2021-09/2021 and 02/2022-07/2022), UPEC, Créteil
Aria Vahidi, Master intern (03/2020-08/2020), Centrale Lille, Lille
• Jobs
Looking for
enthusiastic and creative Master, PhD, and Postdoc
candidates with an outstanding background in artificial intelligence,
image processing, molecular dynamics simulations, or related fields, wishing
to work on projects on the frontiers of molecular biosciences and software
engineering. Interested candidates should contact us with a cover letter, CV, and email
addresses of 1-2 references.
• Research highlights
We have developed ContinuousFlex [1], a software package containing methods for extracting information on continuous conformational
flexibility of biomolecular
complexes from single particle cryo-EM images and
cryo-ET subtomogram.
The open-source code and the installation instructions can be found at GitHub and PyPI. It is also available as a plugin for Scipion.
Methods available in ContinuousFlex:
- HEMNMA: Hybrid Electron Microscopy Normal Mode Analysis
method to interpret heterogeneity of a set of single particle cryo-EM images in terms of continuous
macromolecular conformational transitions, based on normal mode analysis [2-4]
- StructMap: Structural Mapping method to interpret
heterogeneity of a set of single particle cryo-EM
maps in terms of continuous conformational transitions, based on normal mode analysis [5]
- HEMNMA-3D: Extension of HEMNMA to continuous conformational variability analysis of macromolecules in cryo-ET subtomograms (in vitro and in situ) [6]
- TomoFlow: Method for analyzing continuous conformational variability of macromolecules in cryo-ET subtomograms (in vitro and in situ) based on 3D dense optical flow [7]
- NMMD: Method for flexible fitting of atomic structures into cryo-EM maps using a combination of Normal Mode (NM) analysis and Molecular Dynamics (MD) simulations implemented in GENESIS [8]
- DeepHEMNMA: A deep learning extension of HEMNMA [9]
- MDSPACE: Method for extracting atomic-resolution landscapes of continuous conformational variability of biomolecules from cryo-EM single particle images based on a new 3D-to-2D flexible fitting method, which uses Molecular Dynamics (MD) simulation in combination with normal modes, and is embedded in an iterative conformational-landscape refinement scheme [10,15]
- Data synthetis tools: HEMNMA and HEMNMA-3D protocols additionally provide tools for synthesizing noisy and CTF-affected single particle cryo-EM images (HEMNMA) and noisy, CTF and missing wedge affected cryo-ET subtomograms (HEMNMA-3D) with flexible or rigid biomolecular conformations, for several types of conformational distributions, from a given atomic structure or an EM map. One part of the noise is applied on the ideal projections before and the other after the CTF, as described in [11-12].
- MDTOMO: Method for extracting atomic-resolution landscapes of continuous conformational variability of biomolecules from cryo-ET subtomograms, which uses Molecular Dynamics (MD) simulation in combination with normal modes, and is embedded in an iterative conformational-landscape refinement scheme [13,15]
Applications of ContinuousFlex methods:
- Nucleosome studies by in situ cryo-ET [6,7]
- Yeast 70S ribosomes studies by single-particle cryo-EM [9,10]
- SARS-CoV-2 spike studies by cryo-ET [13]
- ATPase p97 studies by single-particle cryo-EM [16]
- HER2-trastuzumab-pertuzumab studies by single-particle cryo-EM [17]
Online lectures:
- Shapes seminar series, Paris,
Feb 7th, 2024 [YouTube]
-
From cryo-EM data to continuous conformational landscapes
- ARC Centre for Cryo-EM of Membrane Proteins, Melbourne, Australia,
November 4th, 2022 [YouTube]
-
DeepHEMNMA for analyzing continuous conformational heterogeneity in single-particle cryo-EM images
- Instruct Image Processing Center (I2PC) seminar series,
April 15th, 2021 [YouTube]
-
HEMNMA and HEMNMA-3D for
in vitro and in situ studies
of continuous conformational variability of macromolecular
complexes, ContinuousFlex plugin of Scipion
- One World Cryo-EM seminar series, March 24th, 2021 [YouTube]
- Combining normal mode analysis,
image analysis, and deep learning for in vitro and in
situ studies of continuous conformational variability of macromolecular
complexes
Selected publications:
[1] Harastani M, Vuillemot R, Hamitouche I, Moghadam NB, Jonic S: ContinuousFlex: Software package for analyzing continuous conformational variability of macromolecules in cryo electron microscopy and tomography data. J Struct Biol. 2022;214:107906. [Journal][Author’s version]
[2] Jin Q, Sorzano CO,
de la Rosa-Trevin JM, Bilbao-Castro JR,
Nunez-Ramirez R, Llorca O, Tama F, Jonic S: Iterative elastic 3D-to-2D alignment
method using normal modes for studying structural dynamics of large
macromolecular complexes. Structure 2014, 22:496-506. [Open-access]
[3] Jonic S: Computational
methods for analyzing conformational variability of macromolecular
complexes from cryo-electron microscopy images.
Curr Opin Struct Biol 2017, 43:114-121. [Journal] [Author’s version]
[4] Harastani M, Sorzano CO, Jonic S: Hybrid Electron Microscopy Normal Mode
Analysis with Scipion. Protein Sci 2020,
29:223-36. [Open-access]
[5] Sanchez Sorzano CO, Alvarez-Cabrera
AL, Kazemi M, Carazo JM, Jonic
S: StructMap:
Elastic Distance Analysis of Electron Microscopy Maps for Studying
Conformational Changes. Biophys J 2016, 110:1753-1765.
[Open-access]
[6] Harastani M, Eltsov M, Leforestier A, Jonic S: HEMNMA-3D:
Cryo Electron Tomography Method Based on Normal
Mode Analysis to Study Continuous Conformational Variability of
Macromolecular Complexes. Front Mol Biosci
2021,8:663121. [Open-access]
[7] Harastani M, Eltsov M, Leforestier A, Jonic S: TomoFlow: Analysis of continuous conformational variability of macromolecules in cryogenic subtomograms based on 3D dense optical flow. J Mol Biol 2021, 167381. [Journal] [Author’s version]
[8] Vuillemot R, Miyashita O, Tama F, Rouiller I, Jonic S: NMMD: Efficient Cryo-EM Flexible Fitting Based on Simultaneous Normal Mode and Molecular Dynamics atomic displacements. J Mol Biol 2022, 167483. [Journal] [Author’s version]
[9] Hamitouche I and Jonic S: DeepHEMNMA: ResNet-based hybrid analysis of continuous conformational heterogeneity in cryo-EM single particle images. Front Mol Biosci 2022 9:965645.[Open-access]
[10] Vuillemot R, Mirzaei A, Harastani M, Hamitouche I, Fréchin L, Klaholz BP, Miyashita O, Tama F, Rouiller I, Jonic S: MDSPACE: Extracting continuous conformational landscapes from cryo-EM single particle datasets using 3D-to-2D flexible fitting based on Molecular Dynamics simulation. J Mol Biol 2023,167951. [Journal][Author’s version]
[11] C.O.S. Sorzano, S. Jonic, R. Núñez-Ramírez, N. Boisset, J.M. Carazo: Fast, robust, and accurate determination of transmission electron microscopy contrast transfer function. J Struct Biol. 2007, 160: 249-262. [Journal]
[12] Jonic S, Sorzano CO, Thevenaz P, El-Bez C, De Carlo S, Unser M: Spline-based
image-to-volume registration for three-dimensional electron microscopy.
Ultramicroscopy
2005, 103:303-317. [Journal]
[13] Vuillemot R, Rouiller I, Jonic S: MDTOMO method for continuous conformational variability analysis in cryo electron subtomograms based on molecular dynamics simulations. Sci Rep 2023, 13, 10596. [Open-access]
[14] Harastani M and Jonic S: Methods for analyzing continuous conformational variability of biomolecules in cryo electron subtomograms: HEMNMA-3D vs. traditional classification. BioRxiv 2021 [Open-access]
[15] Vuillemot R, Harastani M, Hamitouche I, Jonic S : MDSPACE and MDTOMO Software for Extracting Continuous Conformational Landscapes from Datasets of Single Particle Images and Subtomograms Based on Molecular Dynamics Simulations: Latest Developments in ContinuousFlex Software Package. Int J Mol Sci 2024. 25, 20 [Open-access]
[16] Valimehr S, Vuillemot R, Kazemi M, Jonic S, Rouiller I: Analysis of the Conformational Landscape of the N-Domains of the AAA ATPase p97: Disentangling Continuous Conformational Variability of Partially Symmetrical Complexes. Int J Mol Sci 2024, 25, 3371. (2 co-first authors) [Open-access]
[17] Ruedas R, Vuillemot R, Tubiana T, Winter JM, Pieri L, Arteni AA, Samson C, Jonic S, Mathieu M, Bressanelli S: Structure and conformational variability of the HER2-trastuzumab-pertuzumab complex . J Struct Biol 2024, 216(2): 108095 [Journal][Author’s version]
Published datasets (open-access):
[1] Vuillemot R & Jonic S (2022) Data used in J Mol Biol 167951, 2023 https://doi.org/10.5281/zenodo.7415104
[2] Hamitouche I & Jonic S (2022) Data used in Front Mol Biosci 9: 965645, 2022. https://doi.org/10.5281/zenodo.7051222
[3] Harastani M & Jonic S (2021) Data used in J Mol Biol 434(2):167381, 2022. https://doi.org/10.5281/zenodo.5718820
[4] Harastani M, Eltsov M, Leforestier A, Jonic S (2021) Data used in Front Mol Biosci 8:663121, 2021 https://www.ebi.ac.uk/emdb/EMD-12699 and https://doi.org/10.6019/empiar-10679